Wheaton Wood
A Genetic & Genealogical Forest
LESSON 6: atDNA SNPs & Segments
In the last lesson I stated an atDNA test looks at areas of our DNA where we tend to have differences. I want to underscore this because it is something that confuses people. Reporting the sequence of the whole human genome is costly. Since so much of the human genome is identical, it is more practical to do “spot checks” at various places where SNPs (mutations) are known to have occurred. Most of the time these SNPs are separated by a few thousand base pairs before you reach the next tested SNP but we often act as if they are adjacent. SNPs have the highest variability, between individuals and groups. All the base pairs in between vary little, or not at all, even between vastly different people.
Think of your DNA as 22 streets with the names Chromosome 1, Chromosome 2 etc. Some of the streets are longer than others (Chromosome 1 has many more addresses than Chromosome 22). Each street has houses on both sides of the street and in our example each pair of houses facing each other have the same address (shown in the blue box as one address–in essence one nucleotide see Lesson 5). One side of the street represents the values you inherited from Mom and the other side is what you got from Dad. These are expressed as A, G, C or T that denote one of the 4 bases. We have many, many houses that are identical and we just skip over these addresses moving quickly down the street and we stop and take a closer look at the houses where there is variation (a SNP has occurred). Rather than test all these houses that are the same we leave them out and just test where there are SNPS.
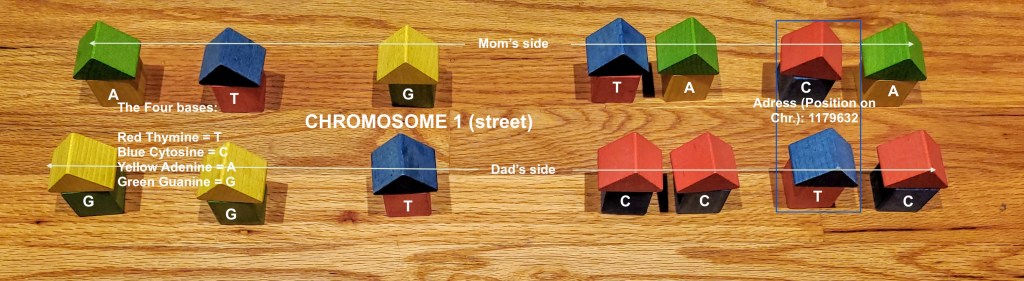
So in this photo we have a street called Chromosome 1. I have put a box around the address 1179632. We have a “C” on Mom’s side of the street and a “T” on Dad’s side. These would appear on a readout or RAW DNA file as:
1179632 CT
PHASING
In a readout it could read “CT” or “TC” and we would not know which parent the “C” or “T” comes from. Here is where “phasing” comes into our example. Phasing simply separates the houses into Mom’s side and Dad’s side. The results from Ancestry uses a special program that attempts to “phase” or “pseudo phase” the SNPs into Mom’s and Dad’s side of the street. Usually this works very well but occasionally this can have the unfortunate effect of making a “true match” look like a non-match by misattributing some of the values to the wrong parent. In the case of misattribution, a series of houses may be on (Dad’s side) but actually be from mom.
There are several other ways we can phase DNA. We can use a program that attempts to separate mom and dad’s contribution. Or if we are lucky enough to have mom and dad and a child we can compare the values that each parent contributes to the child and actually phase what came from where.
SEGMENTS
Most people do not realize that when we have a DNA matching segment with another person we only match half of our DNA with our match (HIR). Going back to our street analogy a match represents a series of Houses (SNPS) only on one side of the street. Only one of the pair of houses at each SNP address needs to match yours. So if your SNP is CT as in our photo example at address 1179632. And your match has TG that is a match for that SNP. In a segment it would be a series of SNPS and only one of the two values for each address needs to match. The only time we are likely to encounter fully identical segments are in children who inherited the same segment from each parent for a given portion of a chromosome. Since the parents each have DNA from their two parents, when those parents give DNA to their children it can be from the same or different grandparents of the child in any given segment as long as one half comes from mom and one half comes from dad.
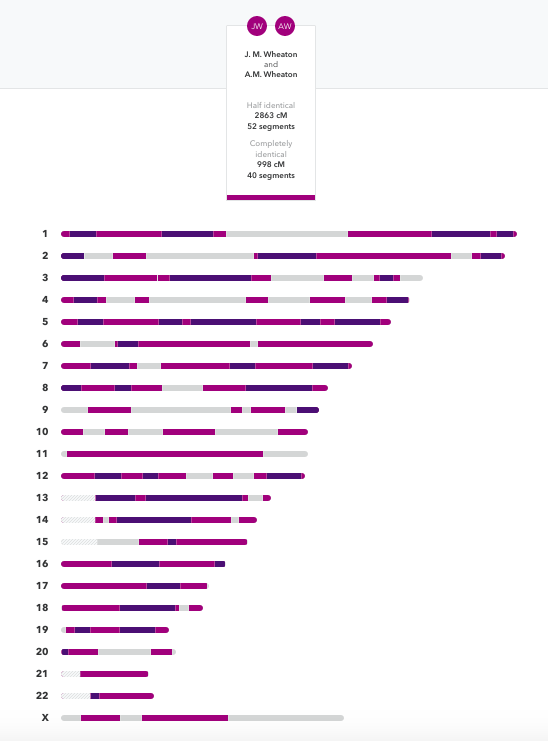
Below is a screenshot from 23andme comparing two of my children. The gray areas on the chromosome is where they each inherited a different segment from each of their parents. For example: A.M. Wheaton may have gotten my mother’s segment from me (maternal grandmother) and J. Wheaton my father’s segment (maternal grandfather). Remember each pair of houses represent SNPS from mom and dad. From their dad A.M. Wheaton got his dad’s dad segment (paternal grandfather) and J. Wheaton from his dad’s mother’s segment (paternal grandmother). So in these gray stretches all 4 grandparents are represented because each child’s two sides do not match the other child’s two sides! In the light purple segments one half of the DNA is the same, so they each may have inherited perhaps my mother’s segment (maternal grandmother) but received different segments from their dad’s parent’s (one inherited the paternal grandmother and the other the paternal grandfather). And finally the dark purple segments show where they got exactly the same segments from both parents. So perhaps my dad’s segment from me (maternal grandfather) and their dad’s dad from him (paternal grandfather).
Remember each child receives half of their DNA from each parent who in turn got half from each parent. All children receive “approximately” 25% of their DNA from each of their four grandparents. The difference between siblings is they each get a different scrambled mix of segments. Have a look at the Visual DNA chart from the first lesson. And just imagine that the patterns of colored bars are randomly different. That’s why children of the same set of parents are so different.
CENTIMORGANS
Each chromosome has a different distribution of SNPs; some are very SNP rich and some not so much. So in our analogy one stretch of street we may need to go many houses or base pairs until we hit a SNP and the other chromosome may have several SNPS on one block. In order to be able to compare the relevance of a segment of DNA we have a unit of measurement that takes into account how many SNPs that different segments contain and how likely a stretch of DNA denotes a likelihood of being genealogically significant. The measurement is called a centimorgan (we use the abbreviation “cM” so as not to be confused with centimeters “cm”). So a cM represents a stretch of DNA where we have left out all the blocks that tend to be the same and only tested those where SNPs occur. What’s left is a string of SNPs.
When you have grasped these concepts then you are ready to use your DNA for genealogical purposes and actually understand what you are doing! Yay! Don’t be discouraged if you need to read this over a few times and occasionally refer back to it.
ADDITIONAL RESOURCES
How many genomic blocks do you share with a cousin? by the Coop Lab Population and Evolutionary Genetics UC Davis
How many genomic blocks do you share with a cousin? by the Coop Lab Population and Evolutionary Genetics UC Davis
How Do DNA Segments get Smaller by Blaine Bettinger
Kelly Wheaton copyright 2020. All Rights Reserved